The contemplation of crossbreeding with Bos taurus breeds in a Bos indicus population is based on the initial presumption that sufficient additive genetic difference (A) exists between the local and exotic breeds, and/or sufficient heterosis (H) is exhibited in crosses between them, that some form of crossbred animal will be more productive than the local breed. Unless these additive and heterotic effects can be accurately estimated in advance, there is great difficulty in deciding what the appropriate breeding strategy should be. Depending on the absolute and relative values of A and H, the best strategy may be any of the following: breed replacement, sane form of synthetic, rotational crossing, or grading up to half or three-quarter exotic.
It could be enormously expensive for a country to discover the correct strategy by trial and error. The time lost in pursuing inappropriate strategies could run into decades. The delay in achieving possible increases in productivity could be very serious economically. The scale of some animal populations, and the ease with which inappropriate crossbreeding schemes can be introduced via A.I. mean that very widespread disappointment, confusion and economic loss could result from unguided crossbreeding.
All of these considerations serve to emphasise strongly the necessity for well planned trials at the beginning of such a crossbreeding program, to provide an adequate information base on which to design the subsequent crossbreeding strategy. Considerable care and investment is justified in the design and conduct of these trials, because of the scale, duration and economic impact of the breeding programs which follow.
The primary purpose of any such crossbreeding trial is the estimation of A and H with sufficient accuracy and precision for subsequent plans to be developed with reasonable confidence. If such trials are to be conducted within the first two generations of crossing between the two breeds, they can involve any or all of six generation groups: the two parental breeds, the F1 the F2, and the backcrosses to the two parental breeds. These six groups are as follows:
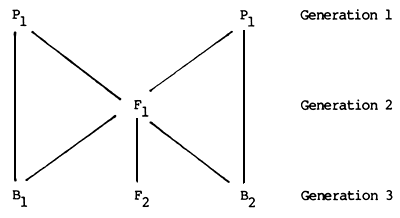
A and H can be estimated from the differences between these groups. It is assumed, of course, that the trial is conducted in such a way that the differences between the groups are not a reflection of environmental, time, location, nutritional or other non-genetic factors. In order to obtain estimates of both A and H, a minimum of three of these groups is required in the trial. Starting with the local population (P1) it is relatively easy to generate F1 offspring using imported semen or males. It may also be possible to provide some animals of the exotic breed (P2) for evaluation in the same environment. This combination o* three groups (both parents and F1) is the most efficient set out of the six possible groups which could be used.
With this optimal set, what size of experiment is required to give an acceptable level of precision in the estimation of A and H. This is the basic question in the design of such trials. Precision is best measured as the standard error of the estimate of A or H. If, for example, the additive difference A is expected to be about 40% of the mid-parent mean, an estimate of this with a standard error equal to 10% of the mid-parent mean (i.e., one-quarter of the actual value estimated) might be regarded as adequate precision for the use of the estimate with confidence in the development of breeding plans. Similarly, if H were expected to be approximately 20% of mid-parent value, then a standard error of 5% (of mid-parent value) might be regarded as adequate precision. As the scale of the experiment goes up, the size of these standard errors of A and H comes down. It is then a matter of judgement as to what balance of precision versus scale is acceptable.
Table 6.1 shows the scale of experiment (with two parental groups and F1) required for given levels of precision for the estimation of A and H. Traits will differ in their inherent variability, and this in turn will affect, the relationship between precision and scale. Three levels of basic variability are therefore provided for: coefficients of variation of 25, 35, and 45%. To achieve given levels of the standard error of A or H (2.5, 5, 10, 15, 20%) the number of animals required in the trial is indicated. In all cases, optimal allocation of numbers to the three groups is assumed.
TABLE 6.1.
NUMBERS OF ANIMALS REQUIRED TO GIVE
SPECIFIED) STANDARD ERRORS (SF) OF A OR H AT DIFFERENT LEVELS OF VARIATION. (SE
AND CV BOTH MEASURED AS % OF MID PARENT MEAN)
| Standard Error of A or H | Coefficient of Variation | |||||
| 25 | 35 | 45 | ||||
| A | H | A | H | A | H | |
| 2.5 | 579 | 468 | 1135 | 918 | 1876 | 1517 |
| 5 | 145 | 117 | 284 | 229 | 469 | 379 |
| 10 | 36 | 29 | 71 | 57 | 117 | 95 |
| 15 | 16 | 13 | 32 | 25 | 52 | 42 |
| 20 | 9 | 7 | 18 | 14 | 29 | 24 |
The following example illustrates the use of the table. If the main trait of interest has a coefficient of variation of 35%, and the standard errors of A and H are each required to be no greater than 5%, then the experiment should contain 284 animals to give this level of precision for the estimation of A, while 229 animals will achieve the desired precision in the estimation of H. Because the design is the same in all cases, H is always more precisely estimated than A (about 20% fewer animals being required to give the same precision).
In these calculations, an optimum allocation of animals over the three groups is assumed. This is defined as an allocation of the total number of animals available over the three groups in such a way as to minimise the sum of the variances of A and H. In the case of this particular design (two parents and F1) the optimum is achieved by allocating 34.5% of the animals respectively to the two parental groups, and 31% to the F1 group. In the example given above, therefore, the 284 animals in the experiment would be allocated 98 each to the two parental groups, and 88 to the F1 group.
The optimal set discussed above includes only the first two generations (both parents and F1). In the next generation, three groups are possible: F2 obtained by inter se mating of F1; B1 obtained by backcrossing F1 to parent I; B2 obtained by backcrossing F1, to parent 2. There can be difficulties in having these three groups comparable to the parental and F1 groups, because they are generated at a later point in time. However, it may be possible to generate further samples of the parental and F1 groups to give valid comparisons.
Assuming that problems of this nature can be overcome, and that all three of these additional groups can be made available, do they contribute to the value of the experiment? One way to respond to this question is to specify a fixed total number of animals in the trial, and to reallocate a certain proportion of them from the parent and F1, groups to the F2 and backcross groups. We can then observe the effect on the actual standard errors of A and H obtained. The results of this calculation are given in Table 6.2.
TABLE 6.2.
THE EFFECT ON THE STANDARD ERRORS OF A & H OF REALLOCATING
RESOURCES FROM PARENTAL AND Fl GROUPS TO BACKCROSSES AND F2 GROUPS.
| Percent of animals reallocated from P1, P2 & F1 to B1, B2 & F2 groups | Relative size of Standard Errors of | |
| A | H | |
| 0 | 100 | 100 |
| 10 | 104 | 105 |
| 20 | 109 | 110 |
| 30 | 114 | 117 |
| 40 | 120 | 125 |
| 50 | 127 | 136 |
It can be seen that for fixed total experimental resources, the inclusion of these extra three groups in all cases reduces the precision of the estimates of A and H. If half of the animals are reallocated, the standard errors of the resulting estimates of A and H are increased by 27% and 36% respectively.
In the design of such experiments, it is not always possible to choose the best combination of groups (P1, P2, F1). For example, where P2 is an exotic breed, it may not be possible to include it for practical or financial reasons. However, semen can be readily imported, so that F1 progeny are usually easy to produce. From the F1 generation, it is of course easy to produce F2. Backcrosses to the exotic breed can be generated by further semen importations, while backcrosses to the local breed can be produced either by mating F1 cows to bulls of the local breed, or vice versa.
Table 6.3 shows the effect on the precision of estimation of A and H of using different combinations of the six possible breeding groups in the experiment. For each design, optimal allocation is again assumed, i.e., a distribution of animals over the groups involved in such a way as to minimise the sum of the variances of A and H. The final column gives the relative scale of experiment (i.e. number of total animals) required to give precision equal to that obtainable with the optimal design.
It can be seen that in all cases the optimal combination (P1, P2, F1) is considerably more efficient than any other design. The next best design requires at least twice the resources to give the same precision.
TABLE 6.3.
COMPARISON OF THE PRECISION ATTAINABLE WITH DIFFERENT COMBINATIONS
OF P1, P2, F1, B1, B2 AND F2 GROUPS. OPTIMAL ALLOCATION TO GROUPS MINIMISES
V(A) + V(H).
| Optimal percent of total animals allocated to groups | Relative size of Standard errors of | Relative number of animal needed for equal precision | ||||||
| P1 | P2 | F1 | B1 | F2 | B2 | A | H | |
| 35 | 35 | 31 | - | - | - | 100 | 100 | 100 |
| 37 | 23 | - | 41 | - | - | 111 | 191 | 231 |
| 29 | 29 | - | - | 41 | - | 109 | 188 | 223 |
| 23 | 37 | - | - | - | 41 | 111 | 191 | 231 |
| 22 | - | 30 | - | 48 | - | 335 | 214 | 826 |
| 22 | - | 35 | - | - | 43 | 170 | 145 | 254 |
| 17 | - | - | 47 | 37 | - | 366 | 385 | 1405 |
| 26 | - | - | 46 | - | 29 | 198 | 288 | 587 |
| 13 | - | - | - | 49 | 38 | 360 | 558 | 2111 |
| - | - | 19 | 38 | 43 | - | 370 | 254 | 1047 |
| - | - | 31 | 35 | - | 35 | 200 | 200 | 400 |
| - | - | 19 | - | 43 | 38 | 370 | 254 | 1047 |
| 28 | 28 | - | 22 | - | 22 | 102 | 186 | 212 |
| 27 | 31 | 30 | 13 | 108 | 187 | 221 | ||
ACKNOWLEDGEMENT
The collaboration of Dr. John Connolly in Chapter 6 is acknowledged.